Abstract
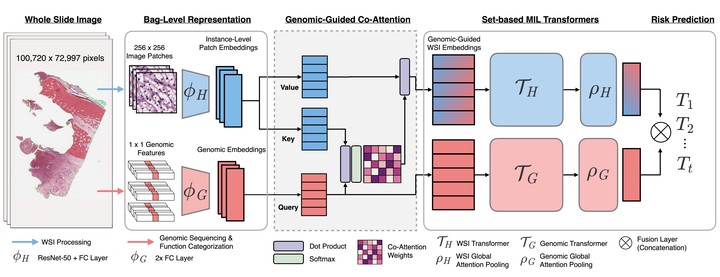
Survival outcome prediction is a challenging weakly-supervised and ordinal regression task in computational pathology that involves modeling complex interactions within the tumor microenvironment in gigapixel whole slide images (WSIs). Despite recent progress in formulating WSIs as bags for multiple instance learning (MIL), representation learning of entire WSIs remains an open and challenging problem, especially in overcoming: 1) the computational complexity of feature aggregation in large bags, and 2) the data heterogeneity gap in incorporating biological priors such as genomic measurements. In this work, we present a Multimodal Co-Attention Transformer (MCAT) framework that learns an interpretable, dense co-attention mapping between WSIs and genomic features formulated in an embedding space. Inspired by approaches in Visual Question Answering (VQA) that can attribute how word embeddings attend to salient objects in an image when answering a question, MCAT learns how histology patches attend to genes when predicting patient survival. In addition to visualizing multimodal interactions, our co-attention transformation also reduces the space complexity of WSI bags, which enables the adaptation of Transformer layers as a general encoder backbone in MIL. We apply our proposed method on five different cancer datasets (4,730 WSIs, 67 million patches). Our experimental results demonstrate that the proposed method consistently achieves superior performance compared to the state-of-the-art methods.